GBS analysis without a reference genome
GBS (Genotyping by Sequencing) utilizes NGS (Next Generation Sequencing) methods similar to Whole Genome Sequencing (WGS). Unlike WGS, which targets the entire genome, GBS involves creating a library from genomic DNA digested by a restriction enzyme. This approach specifically sequences fragments of defined sizes (200-500 bp), making it more cost-effective than other sequencing platforms like WGS.
When a reference genome for the target species is unavailable, a draft genome construction of the genomic region or transcriptome through de novo assembly becomes essential to enhance mapping and genotyping accuracy. By aligning GBS (Genotyping by Sequencing) reads against this assembled draft genome, we can effectively detect SNP (Single Nucleotide Polymorphisms) and small Insertions/Deletions(less than 50 bp).
When a reference genome for the target species is unavailable, a draft genome construction of the genomic region or transcriptome through de novo assembly becomes essential to enhance mapping and genotyping accuracy. By aligning GBS (Genotyping by Sequencing) reads against this assembled draft genome, we can effectively detect SNP (Single Nucleotide Polymorphisms) and small Insertions/Deletions(less than 50 bp).
Work Flow
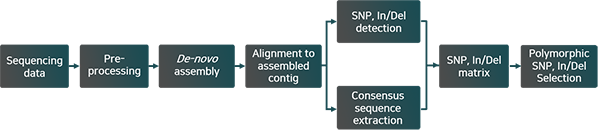
Result Examples
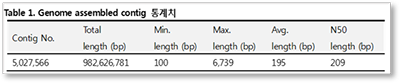 An example of statistics of draft genome assembly
An example of statistics of draft genome assembly
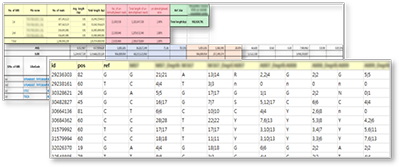 An example of SNP matrix displayed in Excel file format
An example of SNP matrix displayed in Excel file format
Total of 12
related analysis cases
- Analysis of Genetic Relationships and Origin-Specific SNP Marker Exploration in Mauremys reevesii Using GBS Data
- SNP Analysis in Pungitius sinensis Using GBS Data
- Exploration of Markers for Flowering Trait in Persimmons Using GBS Data
- Construction of a Genetic Map and Pseudomolecule Development in Laodelphax striatellus F2 cross population Using GBS Data
- Analysis of Regional Genetic Relationships in Laodelphax striatellus Using GBS Data
- Development of Phylogenetic Trees and Variety-Specific SNP Marker Sets in Dendrobium moniliforme Using GBS Data
- Analysis of Genetic Relationships in Panax ginseng Using GBS Data
- Construction of a Genetic Map and QTL Analysis in Miscanthus sinensis Reciprocal Groups Using GBS Data
- Exploration of Variety-Specific SNP Markers in Colletotrichum Using GBS Data
- Exploration of Variety-Specific SNP Markers in Lilium genetic resources Using GBS Data
- Analysis of Genetic Relationships and Development of Variety-Specific SNP Marker Sets in Freesia refracta Using GBS Data
- Development of Phylogenetic Trees and Variety-Specific SNP Marker Sets in Stevia rebaudiana Using GBS Data
Total of 1
related publication
Total of 11


